IN 2010, when the European Union (EU) passed a directive to eliminate the use of animals in scientific research – aptamers came to the fore to replace animal-derived antibodies that were largely used for developing diagnostic tools.
Aptamers are synthetic antibodies produced without harming a single animal. Aptamers can recognise and bind to their target molecules with high selectivity and affinity. In fact, aptamers, turned out to be a better alternative than animal antibodies – a serendipitous discovery, so to speak.
Aptamers have a more stable nature and a pointed degree of accuracy compared to animal-derived antibodies. Biogenes Technologies – a homegrown technology company established in 2015, developed an aptamer-based Covid-19 test kit as its flagship product.
The company is developing other aptamer-based diagnostic tools using biosensors. These tools are useful for screening sepsis infections, cancer biomarkers, and other non-communicable diseases.
Biogenes has now embarked on aptamer-based tool kits for detecting Human Papillomavirus (HPV) and breast cancer markers.
Tang Kok Mun, the Chief Executive Officer (CEO) of Biogenes says aptamer technology has eased and speeded up the work of scientists in wet laboratories. Before the advent of this technology the cost in producing diagnostic tools was very high, animal-ethical issues and the need for sophisticated refrigeration for the preservation of the antibodies.
However, not everything about aptamers is smooth sailing. Conventional method known as Systemic Evolution of Ligands by Exponential enrichment (SELEX) is time consuming and not highly refined.
Thus, Biogenes took the leap of science to develop Aptamer Computer Aided Design (APTCAD), a web-based computational software which can design aptamers 20 times faster. The software allows users to design aptamers in an integrated and convenient way in two months.
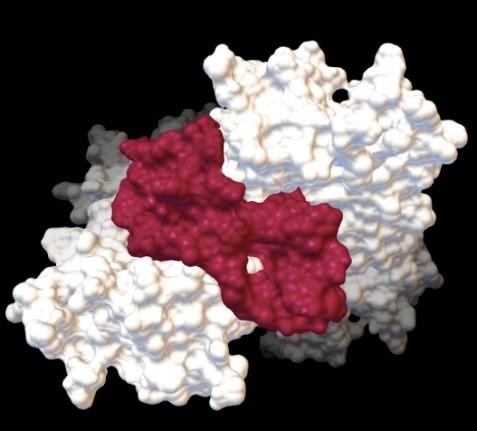
Teffanie Arputheraj, an aptamer expert in Biogenes Technologies, elaborated that, using the APTCAD software, aptamers fold into fixed 3D structures that attach and bind strongly to the target molecules as desired from long chains of nucleotides.
“The conventional method of designing aptamers via SELEX is done by incubating the target molecule in a random library of oligonucleotides, then removing the unbounded oligonucleotides to elute the target bound oligonucleotides. Finally, the eluted sequence is amplified using PCR to observe the binding affinity of the aptamer,” she said.
However, the APTCAD software has surpassed all these steps and generates the 3D structure based on the target molecule nucleotide sequence that is uploaded in the software.
The software calculates the best orientation for these synthetic antibodies to attach strongly to the target molecules. Once, the optimal aptamer sequence is determined via APTCAD, the aptamer can be synthesised and validated in the lab. The validation of aptamer is done using the ELISA protocol.
“A high-speed gaming computer would be faster in generating 3D structure of aptamer compared to a laptop,” adds Teffanie.
“We are still working on the business model for the software but most likely it will be in subscription mode and we are looking forward to training the research community to design aptamer design using this software to strengthen their understanding,” says Tang.
Biogenes Technologies also aspires to digitalise biotechnology as they foresee scientific research to be integrated globally to support swift knowledge sharing. High-affinity aptamers can be uploaded to a global database and someone from a different part of the world can download and validate its efficacy in their wet lab almost immediately. This is in contrast to conventional antibodies that involve material transfer agreement, physical shipping of molecules, and of course cold supply chain and not to mention time and cost.
Moving forward, Biogenes believes that these technological advancements would bring more breakthroughs in the scientific world and break the barriers that are blocking cross-cutting research.
Article originally published in: https://thepetridish.my/2022/10/17/biogenes-leads-the-way-in-aptamer-based-antibodies/